Due to the exponentially growing computational power - sketched by “Moore’s law” - the frontiers of molecular modelling and simulation could be expanded to successively higher levels of theory as well as to a constantly enlarged size of the chemical entities and ensembles under investigation -.ĭissipative particle dynamics (DPD) in particular is a well-established mesoscopic simulation technique to study the structure, dynamics and properties of very large molecular ensembles which may represent millions of atoms. Whereas the fundamental laws of nature are known in principle for nearly a century, their practical application required the development of sufficiently fast computing devices in combination with corresponding theoretical approximations - a venture that successfully forged ahead in the last decades as indicated by the 19 Nobel Prizes in chemistry. Molecular modelling and simulation aims at (at least) theoretically explaining and (at best) predicting the structures, properties and dynamics of molecules and molecular ensembles. Molecular Fragment Cheminformatics may be regarded as a crucial accelerator to propagate MFD and similar mesoscopic simulation techniques in the molecular sciences. Mesoscopic simulation techniques like MFD may be considerably alleviated and broadened for practical use with a consequent cheminformatics layer that successfully tackles its setup subtleties and conceptual usage hurdles.
It is shown that the requirements of the roadmap may be partly covered by already existing open-source cheminformatics software. The fSMILES notation and the following concepts and methods for building blocks 2-4 are outlined with examples and practical usage scenarios. building block 1) is a SMILES-like line notation (denoted fSMILES) with connected molecular fragments to represent a molecular structure. The basis of the cheminformatics approach (i.e. The molecular fragment cheminformatics roadmap consists of four consecutive building blocks: An adequate fragment structure representation (1), defined operations on these fragment structures (2), the description of compartments with defined compositions and structural alignments (3), and the graphical setup and analysis of a whole simulation box (4). This study aims at outlining a complete cheminformatics roadmap that frames a mesoscopic Molecular Fragment Dynamics (MFD) simulation kernel to allow its efficient use and practical application. A consequent cheminformatics approach tackles these problems and provides solutions for a considerably enhanced accessibility. Therefore a pure kernel-based mesoscopic simulation task is a tedious, time-consuming and error-prone venture that limits its practical use and application.
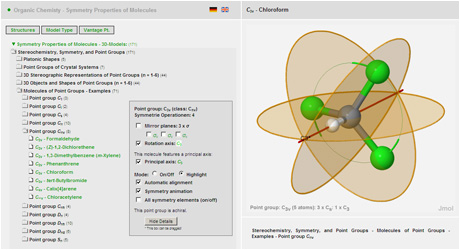
#JMOL SIMULATION SOFTWARE#
For its simulation setup and output a mesoscopic simulation kernel software uses abstract matrix (array) representations for bead topology and connectivity. Mesoscopic simulation studies the structure, dynamics and properties of large molecular ensembles with millions of atoms: Its basic interacting units (beads) are no longer the nuclei and electrons of quantum chemical ab-initio calculations or the atom types of molecular mechanics but molecular fragments, molecules or even larger molecular entities.
